Development
Data computing with Python

In my previous blog article, we learned how to clean data with Python Pandas. Today we are going to learn one of the key tools that has been widely used for data processing and data intensive computing.
Python is one of the most popular languages for data analysis and development, because it’s fast for testing and coding. However, this interpreted, dynamically-typed, high-level language, is somewhat slow for advanced/complex computing tasks.
To get the best of both worlds (fast code development time and fast code execution time): NumPy comes in.
In this article, we’ll be going through some basic NumPy functions and how to use them effectively when working with a large dataset. We will also take a look at the difference between Pure Python and NumPy. Finally, we will learn some strategies that can help us speed up code with NumPy.
Introduction to NumPy
What is NumPy?

NumPy is a Python library used for working with multi-dimensional arrays. NumPy stands for “Numerical Python”. It provides a high-performance multidimensional array object, and tools for working with these arrays.
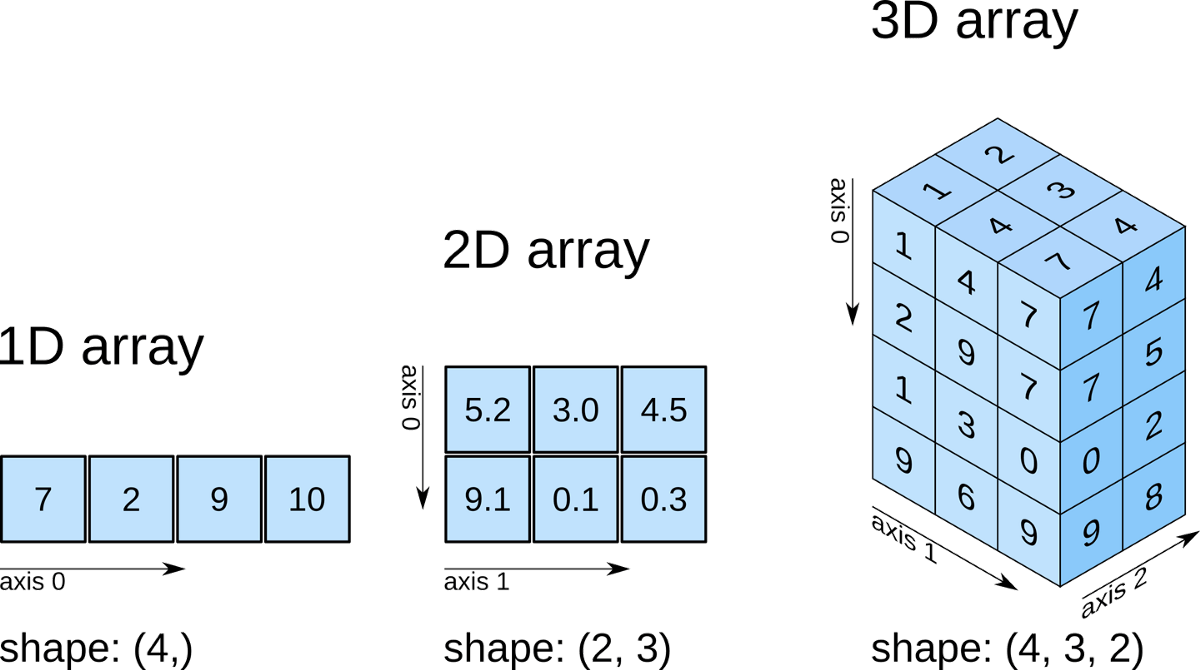
NumPy is a core Python library for scientific computing. With it, we can perform complex mathematical and logical operations faster and easier, such as:
- Vector-Vector multiplication
- Matrix-Matrix and Matrix-Vector multiplication
- Element-wise operations on vectors and matrices (i.e., adding, subtracting, multiplying, and dividing by a number )
- Element-wise or array-wise comparisons
- Applying functions element-wise to a vector/matrix ( like pow, log, and exp)
- A whole lot of Linear Algebra operations can be found in NumPy.linalg
- Reduction, statistics, and much more.
Quick Start to NumPy
Let’s take a look at a few examples to understand it better.
1. Creating Arrays
With Numpy, we can create multi-dimensional arrays:
import numpy as np
# Create a 1-D Array
arr = np.array([1, 2, 3, 4, 5])
print(arr)
# Create a 2-D Array
arr = np.array([[1, 2, 3], [4, 5, 6]])
print(arr)
# Create a 3-D Array
arr = np.array([[[1, 2, 3], [4, 5, 6]], [[1, 2, 3], [4, 5, 6]]])
print(arr)
Out:
array([1, 2, 3, 4, 5])
---
array([[1, 2, 3],
[4, 5, 6]])
---
array([[[1, 2, 3],
[4, 5, 6]],
[[1, 2, 3],
[4, 5, 6]]])
2. Array Indexing
Indexing arrays is similar to Python lists:
arr = np.array([1, 2, 3, 4])
# Get the first element from the array
print(arr[0])
# Get third and fourth elements from the array and sum them.
print(arr[2] + arr[3])
Out:
1
---
7
3. Array Slicing
We can get a slice of an array like this: [start:end].
arr = np.array([1, 2, 3, 4, 5, 6, 7])
# Slice elements from index 1 to index 5 from the array
arr[1:5]
Out:
array([2, 3, 4, 5])
4. Array Shape
The shape of an array is the number of elements in each dimension. We can get the current shape of an array like this:
# Create a 2-D array
arr = np.array([[1, 2, 3, 4], [5, 6, 7, 8]])
print(arr)
# Get the shape
print(arr.shape)
Out:
array([[1, 2, 3, 4],
[5, 6, 7, 8]])
---
(2, 4)
(2, 4) means that the array has two dimensions, the first dimension has two elements and the second has four.
5. Reshaping arrays
Reshape an array from 1-D to 2-D
# Create a 1-D array
arr = np.array([1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12])
print(arr)
# Convert the 1-D array with 12 elements into a 2-D array. The outermost dimension will have 4 arrays, each with 3 elements
new_arr = arr.reshape(4, 3)
print(new_arr)
Out:
array([ 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12])
---
array([[ 1, 2, 3],
[ 4, 5, 6],
[ 7, 8, 9],
[10, 11, 12]])
For more details, see NumPy Documentation.
Python Lists vs. NumPy Arrays
A common question that pops up: what is the real difference between Python lists and NumPy arrays? The simple answer is performance.
Why is NumPy faster?
Python speed suffers from small overhead, such as type checking and reference counting, which builds up for each operation and it could become especially significant in a large loop.
What NumPy does is that it pushes those loop operations down into compiled code, so the type checking only has to happen once for the entire loop therefore they can be done quickly.
Overall, NumPy performs better in the following instances:
- Size - NumPy uses much less memory to store data.
- Performance - NumPy arrays are faster than Python lists.
- Functionality - NumPy has optimized mathematical functions built-in that are convenient to use.
Memory consumption
To allocate a list of 1000 elements, Python takes 48000 bytes of the memory, whereas NumPy only needs 8000 bytes. What a significant improvement!
import numpy as np
import sys
# Declaring a Python list of 1000 elements
ls = range(1000)
print(f"Size of each element of the list: {sys.getsizeof(ls)} bytes.")
print(f"Size of the list: {sys.getsizeof(ls)*len(ls)} bytes.")
# Declaring a Numpy array of 1000 elements
arr = np.arange(1000)
print(f"Size of each element of the array: {arr.itemsize} bytes.")
print(f"Size of the array: {arr.size*arr.itemsize} bytes.")
Out:
Size of each element of the list: 48 bytes.
Size of the list: 48000 bytes.
Size of each element of the array: 8 bytes.
Size of the array: 8000 bytes.
Performance
There’s a vast difference in performance. If we add 5 to each element of a list of 100k values:
# Python way
ls = list(range(100000))
%timeit [val + 5 for val in ls]
# Numpy way
arr = np.array(ls)
%timeit arr + 5
Out:
6.17 ms ± 35.9 µs per loop
---
34.8 µs ± 340 ns per loop
It turned out using about 6 milliseconds (could be varied depending on different machines) for Python to process per loop while NumPy only took 34 microseconds per loop. We have about nearly 200x speed up by taking away the loops in Python.
Strategies for speeding up code with NumPy
Now we’ve learned that NumPy is a powerful tool for working with large arrays. However, it’s not only useful for scientific computing. It can also be handy for some software/services development tasks that involve complex data processing with a large dataset. With pure Python, we would normally loop over the dataset and do it raw and that’s… not so optimal right?
In order to achieve that effectively, let’s see what NumPy is capable of and how we can use it to speed our code, by applying the following strategies. Let’s get rid of those slow loops!
- Ufuncs
- Aggregations
- Slicing, Masking and Fancy indexing
Ufuncs
Ufuncs, short for universal functions, offers a bunch of operators that operate element by element on entire arrays.
(See universal functions basics for more details about ufuncs.)
For example, we want to add the elements of list a and b. Without NumPy, we can use the built-in zip() method:
a = list(range(1001)) # [0, 1, 2, 3, ..., 999, 1000]
b = list(reversed(range(1001))) # [1000, 999, 998, ..., 1, 0]
c = [i+j for i, j in zip(a, b)] # returns [1000, 1000, ..., 1000, 1000]
Instead of looping the list, we can simply use the add() from Numpy ufunc:
import numpy as np
a = np.array(list(range(1001)))
b = np.array(list(reversed(range(1001))))
c = np.add(a, b) #returns array([1000, 1000, ..., 1000, 1000])
Let’s compare the execution time:
%timeit [i+j for i, j in zip(a, b)]
%timeit np.add(a, b)
Out:
61 µs ± 4.11 µs per loop
---
811 ns ± 29.8 ns per loop
We got 75x faster with the NumPy version and the code looks much cleaner. Isn’t that awesome?
There are many other ufuncs available:
- Arithmetic:
+,-,*,/,//,%… - Comparison:
<,>,==,<=,>=,!=… - Bitwise:
&,|,~,^… - Trig family:
np.sin,np.cos,np.tan… - And more…
Aggregations
Aggregations are functions which summarize the values in an array (e.g. min, max, sum, mean, etc.).
Let’s see how long it takes a Python loop to get the minimum value of a list of 100k random values.
With Python built-in min():
from random import random
ls = [random() for i in range(100000)]
%timeit min(ls)
With NumPy aggregate functions min():
arr = np.array(ls)
%timeit arr.min()
Out:
1.3 ms ± 7.59 µs per loop
---
33.6 µs ± 248 ns per loop
Again, we got about 40x speedup without writing a single loop with NumPy!
In addition, aggregations can also work on multi-dimensional arrays:
# Create a 3x5 array with random numbers
arr = np.random.randint(0, 10, (3, 5))
print(arr)
# Get the sum of the entire array
print(arr.sum())
# Get the sum of all the columns in the array
print(arr.sum(axis=0))
# Get the sum of all the rows in the array
print(arr.sum(axis=1))
Out:
array([[7, 7, 5, 1, 9],
[8, 7, 0, 2, 4],
[0, 5, 6, 4, 0]])
---
65
---
array([15, 19, 11, 7, 13])
---
array([29, 21, 15])
There are more aggregate functions:
- sum
- mean
- product
- median
- variance
- argmin
- argmax
- And more
Slicing, Masking and Fancy indexing
We just saw how to slice and index NumPy arrays earlier. In fact, there are other faster and more convenient ways that NumPy offers:
Masking
With masks, we can index an array with another array. For example, we want to index an array a with a boolean mask. What we get out, is that only the elements that line up with True in that mask will be returned:
# Create an array
arr = np.array([1, 2, 3, 4, 5])
print(arr)
# Create a boolean mask
mask = np.array([False, False, True, True, True])
print(arr[mask])
Out:
array([1, 2, 3, 4, 5])
---
array([3, 4, 5])
Here we only get [3 4 5], because these values line up with True in the mask.
Now you might be a little confused and wonder why we would ever need that? Where masks become useful is when they are combined with ufuncs. For example, we want to get the elements from an array based on some conditions. We can index that array using a mask with the conditions:
# Create an array
arr = np.array([1, 2, 3, 4, 5])
print(arr)
# Create a mask
mask = (arr % 2 ==0) | (arr > 4)
print(arr[mask])
Out:
array([1, 2, 3, 4, 5])
---
array([2, 4, 5])
And the values that meet the criteria will be returned.
Fancy indexing
The idea of fancy indexing is very simple. It allows us to access multiple array elements at once by passing an array of indices.
For example, we get the 0th and 1st element from array arr by passing another array of the indices:
arr = np.array([1, 2, 3, 4, 5])
index_to_select = [0, 1]
print(arr[index_to_select])
Out:
array([1, 2])
Let’s combine all these together:
- Access multi-dimensional arrays by
- [row, column]
- :
# Create a 2x3 array
arr = np.arange(6).reshape(2, 3)
print(arr)
# Get row 0 and column 1
print(arr[0, 1])
# Get all rows in column 1 (Mixing slices and indices)
arr[:, 1]
Out:
array([[0, 1, 2],
[3, 4, 5]])
---
1
---
array([1, 4])
- Masking multi-dimensional arrays:
# Create a 2x3 array
arr = np.arange(6).reshape(2, 3)
print(arr)
# Masking the entire array
mask = abs(arr-3)<2
print(arr[mask])
Out:
array([[0, 1, 2],
[3, 4, 5]])
---
array([2, 3, 4])
- Mixing masking and slicing:
arr = np.arange(6).reshape(2, 3)
print(arr)
mask = arr.sum(axis=1) > 4
print(arr[mask, 1:])
Out:
array([[0, 1, 2],
[3, 4, 5]])
---
array([[4, 5]])
All of these NumPy operations can be combined in nearly limitless ways!
Conclusion
No matter how large your dataset is or how complex are the operations you need to perform with your data, NumPy functions are there to make it easier and faster.
Let’s summarize what we’ve covered:
- Writing Python is fast, but loops are slow for large dataset computing.
- NumPy is a great tool to use! It pushes loops into its compiled layer, so we get:
- Faster development time, and
- Faster execution time.
- Strategies for speeding up your code:
- Ufuncs - For element wise operations.
- Aggregations - For array summarization.
- Slicing, masking and fancy indexing - For quickly selecting and operating on arrays.
Looking for the final source code!
References
- Python Course - NumPy Tutorial
- NumPy Documentation

Did this article start to give you some ideas? We’d love to work with you! Get in touch and let’s discover what we can do together.












